Precision Epilepsy How Gene-Targeted Therapies Are Changing Seizure Treatment in 2026
Key Takeaways
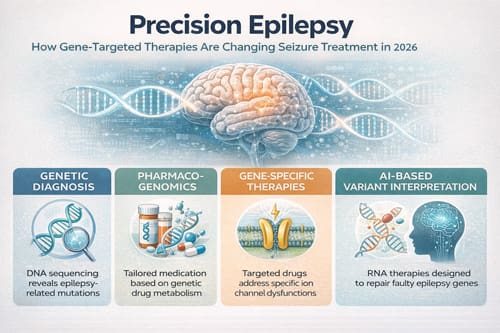
Precision epilepsy is redefining the clinical management of seizure disorders by shifting treatment strategies from generalized symptom suppression toward interventions directed at the underlying molecular and genetic causes of disease. This transition is particularly important for the substantial proportion of patients who remain refractory to conventional therapy, as approximately 30 to 40 percent of individuals with epilepsy continue to experience seizures despite treatment with available antiseizure medications. Although the number of approved antiseizure drugs has expanded considerably over recent decades, the burden of drug resistant epilepsy has remained largely unchanged, highlighting the limitations of traditional trial based prescribing approaches and the need for more biologically informed treatment models.
The growing understanding of epilepsy genetics has revealed that pathogenic variants contribute significantly to epileptogenesis across both pediatric and adult populations. Genetic abnormalities affecting ion channels, neurotransmitter receptors, synaptic regulation, metabolic pathways, and intracellular signaling mechanisms are now recognized as central drivers in many epilepsy syndromes. This burden is substantial on a global scale, with genetic epilepsy affecting millions of individuals, including more than nine million people in China alone. These discoveries have accelerated the emergence of precision medicine approaches that integrate genomic diagnosis, pharmacogenomic risk assessment, and gene specific therapeutic strategies into routine epilepsy care.
One of the most significant developments in this field is the increasing role of genetic testing as a first line diagnostic tool, particularly in patients with early onset epilepsy, developmental epileptic encephalopathies, or unexplained drug resistance. Whole genome sequencing and related genomic technologies now achieve diagnostic yields of approximately 20 to 40 percent, depending on patient selection and phenotype complexity. Importantly, the clinical value of testing extends beyond diagnosis alone. Evidence indicates that genomic findings alter clinical management in nearly half of tested patients, with treatment modification leading to measurable seizure reduction in approximately 65 percent of cases. Identification of a causative variant can influence medication selection, determine eligibility for targeted therapies, refine prognostic counseling, and guide family planning discussions.
Pre treatment pharmacogenomic screening has also become an increasingly important element of safe epilepsy care. Certain antiseizure medications carry significant risks of severe hypersensitivity reactions in genetically susceptible populations. A well established example is the association between carbamazepine and the HLA B*15:02 allele, which is strongly linked to Stevens Johnson syndrome and toxic epidermal necrolysis, particularly among individuals of Asian ancestry. Screening for this allele before treatment initiation has been shown to reduce the risk of severe cutaneous reactions by approximately 25 to 220 fold, making it one of the clearest examples of effective genomic risk prevention in clinical neurology. Similar attention is increasingly being given to other pharmacogenetic markers that affect drug metabolism, efficacy, and toxicity.
The therapeutic impact of precision epilepsy is especially evident in gene specific interventions, where treatment selection is guided by the biological pathway affected by a specific mutation. In tuberous sclerosis complex, where dysregulation of the mammalian target of rapamycin pathway drives epileptogenesis, everolimus has demonstrated approximately 40 percent seizure reduction and has become an important adjunctive therapy in affected patients. In glucose transporter type 1 deficiency syndrome, where impaired glucose transport across the blood brain barrier limits cerebral energy availability, ketogenic diet therapy directly addresses the metabolic defect and achieves seizure freedom in approximately 64 percent of patients. Similarly, highly targeted responses have been observed in rare syndromes such as NR2F1 related epilepsy, where perampanel has produced complete seizure control in reported cases, illustrating how mechanistic alignment between mutation and therapy can dramatically improve outcomes.
Among the most promising emerging developments are antisense oligonucleotide therapies, which offer the ability to directly modify gene expression at the RNA level. These therapies represent a major advance for disorders previously considered untreatable. Zorevunersen, developed for Dravet syndrome associated with SCN1A haploinsufficiency, has shown seizure reductions of approximately 85 percent in early clinical studies, making it the most advanced RNA based therapeutic candidate currently under investigation in epilepsy. By increasing production of functional sodium channel protein, this approach directly targets the molecular basis of disease rather than its downstream manifestations.
Artificial intelligence is also becoming an essential component of precision epilepsy, particularly in the interpretation of complex genomic data. A major challenge in clinical genetics remains the classification of variants of uncertain significance, many of which cannot be confidently linked to disease using conventional methods. Machine learning algorithms are increasingly capable of integrating genomic, structural, and clinical data to improve variant interpretation, achieving approximately 80 percent accuracy in classification and reclassifying nearly 60 percent of previously uncertain variants. These tools are particularly valuable for populations historically underrepresented in genomic databases, where conventional reference data may be limited and diagnostic inequities have persisted.
The convergence of genomic diagnostics, pharmacogenomics, targeted therapies, and computational interpretation is fundamentally changing the epilepsy treatment paradigm. Instead of sequentially testing multiple antiseizure medications with uncertain probability of success, clinicians are increasingly able to align therapy with each patient’s unique molecular profile. This approach has the potential not only to improve seizure control but also to reduce adverse effects, shorten time to effective treatment, and improve long term neurodevelopmental outcomes.
As precision epilepsy continues to evolve, several challenges remain, including equitable access to genetic testing, cost of advanced therapies, interpretation of rare variants, and integration of genomic expertise into everyday neurological practice. Nevertheless, the field is moving steadily toward a future in which epilepsy treatment is no longer defined primarily by symptom suppression, but by targeted intervention based on underlying disease biology. In 2026, precision therapy stands as one of the most transformative developments in epilepsy care, offering a realistic path toward more individualized, effective, and scientifically grounded seizure management.
Understanding Precision Medicine in Epilepsy
What Makes Epilepsy a Target for Gene-Based Treatment
Genetic etiologies drive seizures in up to 40% of children and 23% of adults with epilepsy [1]. This substantial genetic contribution creates a foundation for precision medicine approaches that target specific molecular mechanisms rather than empirically suppressing seizure activity. Twin studies reveal heritability estimates ranging from 25% to 70%, with probandwise concordance rates reaching 0.37 in monozygotic twins compared to lower rates in dizygotic twins [2].
Monogenic epilepsy accounts for only 1% to 2% of cases, typically following autosomal dominant inheritance patterns with mutations in protein-coding sequences [3]. The remaining epilepsies are polygenic, possibly caused by an excess of rare variants in epilepsy-associated genes. More than 750 genes have been reported as associated with epileptic conditions, though only a small percentage likely cause monogenic epilepsy [3]. SCN1A, encoding the voltage-gated sodium channel Nav1.1a, represents the most commonly recognized epileptogenic gene with over 2,000 mutations described [3]. This gene causes 80% of Dravet syndrome cases [3].
From Traditional ASMs to Gene-Targeted Therapies
Current medical therapy for most epilepsies remains imprecise, aimed primarily at empirical seizure reduction rather than targeting specific disease processes [2]. Despite increasing availability of antiseizure medications, the incidence of drug-resistant epilepsy has not decreased [4]. Nearly 30% to 40% of individuals with epilepsy continue to experience uncontrolled seizures, highlighting the need for mechanism-driven solutions [5].
The shift toward precision therapy in epilepsy stems from understanding that genetic changes result in metabolic abnormalities requiring specific supplements or dietary interventions rather than conventional ASMs [6]. Pyridoxine treatment for antiquitin deficiency and ketogenic diet for GLUT1 deficiency syndrome exemplify this concept [4]. In essence, these treatments address root molecular defects instead of downstream electrical hyperexcitability.
Preclinical evidence has led to phase 3 clinical trials, such as everolimus for mTOR pathway inhibition in tuberous sclerosis complex [4]. However, preclinical findings do not always translate into effective treatments, as demonstrated by heterogeneous effects of quinidine in KCNT1-related epilepsy [4].
The Role of Genetic Testing in Modern Epilepsy Care
Genetic testing has transformed from a specialized tool to a first-line diagnostic approach. Whole genome sequencing and exome sequencing are recommended by the National Society of Genetic Counselors and American Epilepsy Society as first-tier tests for individuals with unexplained epilepsy [1]. Overall detection rates approximate 20% to 40% depending on the type of epilepsy, number of genes tested, and whether unaffected family members provide comparative data [3].
A genetic diagnosis resulted in clinical management changes for 208 patients out of 418 tested, representing 49.8% of cases [1]. Among 167 patients with follow-up information available, 125 patients (74.9%) reported positive outcomes, with 108 patients (64.7%) experiencing reduction or elimination of seizures [1]. The most common management changes included adding new medication (21.7%), initiating medication (14.2%), specialist referrals (13.4%), and medication cessation (11.7%) [1].
Diagnostic yield varies by clinical presentation. Gene panels achieve detection rates as high as 54% in epilepsy with structural brain malformations, with overall yields around 19% [5]. Age at seizure onset proves highly predictive, with diagnostic yields reaching 38% for onset before three years compared to 4% for later onset [5].
Pharmacogenomic Testing Before Treatment
Pharmacogenomic testing before drug initiation prevents life-threatening adverse reactions in patients receiving antiseizure medications. Two genes dominate pre-treatment screening protocols: HLA-B variants predict immune-mediated hypersensitivity, while CYP2C9 polymorphisms determine metabolic capacity and toxicity risk.
HLA-B*15:02 and Carbamazepine Hypersensitivity Risk
Carbamazepine ranks among the most prescribed antiepileptic drugs, FDA-approved for epilepsy, bipolar disorder, and trigeminal neuralgia [4]. A minority of treated patients develop cutaneous adverse reactions ranging from mild rash to Stevens-Johnson syndrome (SJS) and toxic epidermal necrolysis (TEN), conditions with mortality rates reaching 10% for SJS and 50% for TEN [7].
The HLA-B15:02 allele predicts severe cutaneous reactions with striking ethnic variation. Prevalence ranges from 10% to 15% in Chinese populations, 2% to 4% in Southeast Asians and Indians, and less than 1% in Japanese and Koreans [8]. Within Han Chinese populations treated with carbamazepine, all 44 patients who developed SJS/TEN carried HLA-B15:02, compared to only 3% of tolerant controls [8]. Meta-analyzes demonstrate odds ratios between 25- and 220-fold depending on ethnicity [9].
Studies in specific populations confirm this association. Among Indian patients in Malaysia, predominantly from Southern India, carriers showed an odds ratio of 33.3 for carbamazepine-induced SJS/TEN [10]. In Han Chinese from northeastern China, 22.9% of SJS/TEN cases carried the allele versus 2% of tolerant controls, yielding an odds ratio of 18.2 [10]. Notably, the allele demonstrates predictive value even in populations with low allele frequency, emphasizing the necessity for testing irrespective of ancestral background [10].
As a result, the FDA mandates screening in genetically at-risk populations before carbamazepine initiation [4]. Patients testing positive should avoid carbamazepine unless benefits clearly outweigh risks [7]. Cost-effectiveness analysis in Asian Australian patients demonstrated an incremental cost-effectiveness ratio of USD 10,523 per quality-adjusted life-year, supporting Medicare subsidization [10].
European populations require attention to HLA-A31:01, present in 2% to 5% of Northern Europeans [9]. This variant increases drug-induced hypersensitivity risk from 3.8% to 26% [11]. Phenytoin carries similar HLA-B15:02 associations, particularly among Thai and Chinese Asian populations, though the evidence remains less robust than for carbamazepine [8].
CYP2C9 Variants and Phenytoin Metabolism
CYP2C9 metabolizes approximately 90% of phenytoin, with genetic polymorphisms determining clearance rates and toxicity susceptibility [1]. Two common variants, CYP2C92 and CYP2C93, encode enzymes with reduced activity [8]. The *2 allele occurs in 10% to 20% of Caucasians but only 1% to 3% of Asians, while *3 appears in less than 10% of most populations and proves extremely rare in African populations [8].
Carriers of reduced-function alleles metabolize phenytoin considerably slower than wild-type individuals, increasing concentration-dependent neurotoxicity risk [1]. In South Indian populations, patients with the *1/3 genotype showed 4.8-fold increased odds of toxicity compared to wild-type carriers [5]. The CYP2C911 variant, found in African populations at frequencies reaching 5.1% among Nigerians, reduces phenytoin metabolic activity by more than 50% [5].
Clinical Pharmacogenetics Implementation Consortium guidelines recommend dose adjustments based on metabolizer status. Extensive metabolizers, representing approximately 91% of the population, require no adjustment [9]. Intermediate metabolizers (roughly 8% of patients) should receive at least a 25% initial maintenance dose reduction, while poor metabolizers need at least a 50% reduction [9]. Continued adjustment relies on therapeutic drug monitoring and patient response.
Clinical Implementation of Pre-Treatment Genetic Screening
The presence of both HLA-B*15:02 and CYP2C9 *2 or 3 variants in the same patient amplifies SJS/TEN risk, contraindicating phenytoin use [12]. Recent evidence suggests CYP2C93 associates with cutaneous adverse events independently of HLA-B genotype [13]. Despite available evidence, pre-treatment CYP2C9 testing remains uncommon, with clinical monitoring and serum drug levels constituting standard practice [1]. The clinical value and cost-effectiveness of CYP2C9 genotyping require formal testing in prospective studies [14].

Gene-Specific Therapies in Clinical Use 
Several precision therapies have moved from experimental protocols into standard clinical practice, each addressing specific genetic defects through mechanism-based interventions. These treatments exemplify precision medicine in epilepsy by targeting molecular pathways disrupted by pathogenic variants.
mTOR Inhibitors for TSC1/TSC2 Mutations
Tuberous sclerosis complex arises from mutations in TSC1 or TSC2 genes, causing mTOR pathway hyperactivation that drives seizure generation and developmental abnormalities [15]. Everolimus gained FDA approval in 2018 for adjunctive treatment of refractory focal seizures in TSC patients over two years of age [16]. The EXIST-3 trial demonstrated seizure reductions of 28% for low-exposure and 40% for high-exposure everolimus compared to placebo, with particular efficacy in children under six years [17]. Responder rates showing at least 50% seizure reduction ranged from 27% to 100% across studies with different follow-up periods [15]. Early mTOR inhibitor use may improve white matter abnormalities and reduce the incidence of drug-resistant epilepsy when initiated soon after birth [17].
Ketogenic Diet for SLC2A1 (GLUT1) Deficiency
Mutations in SLC2A1 impair glucose transport across the blood-brain barrier, creating chronic cerebral energy depletion that ketogenic diet therapies address by providing ketones as alternative fuel [18]. The classic ketogenic diet, providing 87% to 90% of calories from fat in 3:1 to 4:1 ratios, achieves seizure freedom in 64% of children with GLUT1 deficiency syndrome [19]. Studies report seizure control rates exceeding 90% reduction in 80% of patients, with better outcomes when treatment starts early [20]. The modified Atkins diet offers improved compliance with fewer side effects while maintaining therapeutic efficacy [18]. Conversely, blood ketone levels above 2.4 µmol/L should be monitored, as 67% of families report correlation between ketosis levels and seizure control [21].
Pyridoxine Treatment for ALDH7A1 Variants
Pyridoxine-dependent epilepsy results from ALDH7A1 mutations causing neonatal seizures unresponsive to standard medications but controlled with pyridoxine [22]. Current dosing recommendations range from 15 to 30 mg/kg/day, not exceeding 500 mg daily to avoid peripheral neuropathy [7]. Triple therapy combining pyridoxine with arginine supplementation and lysine-restricted diet reduces neurotoxic metabolites and enhances neurodevelopmental outcomes when initiated within the first six months of life [11]. Despite excellent seizure control, 75% of patients experience global developmental delay even with early treatment initiation [22].
Perampanel for NR2F1-Related Epilepsy
NR2F1 pathogenic variants cause epileptic encephalopathy previously resistant to multiple medications [23]. Perampanel, an AMPA receptor antagonist, exhibited dramatic efficacy in six patients with NR2F1-related epilepsy, achieving 100% seizure freedom [23]. The average time to seizure control was 7.33 months at dosages of 0.22 mg/kg per day [23]. Developmental delay and intellectual disability symptoms improved in all patients after seizure control [23].
Quinidine for KCNT1 Gain-of-Function Mutations
KCNT1 mutations cause gain-of-function channel activity that quinidine reversibly blocks [24]. Clinical responses prove variable, with effective outcomes defined as greater than 50% seizure reduction occurring in only 50% of treated patients [25]. Quinidine demonstrates preferential blocking of mutant versus wild-type channels at concentrations of 1 to 10 µM [26]. Nevertheless, gastrointestinal adverse effects limit tolerability, occurring in up to 50% of patients and necessitating treatment discontinuation in some cases [27].
Therapeutic Drug Monitoring and Personalized Dosing
Therapeutic drug monitoring optimizes antiseizure medication dosing through quantitative measurement of plasma concentrations combined with clinical pharmacology principles. This approach proves valuable for guiding treatment adjustments, especially in vulnerable populations including children, pregnant women, and those receiving polytherapy.
Next-Day TDM Results Using Mass Spectrometry
Ultra-performance liquid chromatography coupled with tandem mass spectrometry (UPLC-MS/MS) enables rapid, accurate quantification of newer antiseizure medications [10]. This analytical method follows Clinical Laboratory Standards Institute guideline C62-A for liquid chromatography-mass spectrometry validation [10]. The technology simultaneously detects multiple compounds across several orders of magnitude with limits of quantification below 0.05 μg/mL for all analytes [28]. Intraassay and interassay imprecisions remain under 5% coefficient of variation, while accuracy stays within ±15% bias [28].
Orienting therapeutic ranges established through this methodology include gabapentin (2 to 20 μg/mL), lacosamide (1 to 10 μg/mL), lamotrigine (2.5 to 15 μg/mL), levetiracetam (12 to 46 μg/mL), oxcarbazepine (3 to 35 μg/mL), perampanel (180 to 980 ng/mL), pregabalin (2 to 8 μg/mL), topiramate (5 to 20 μg/mL), and zonisamide (10 to 40 μg/mL) [10]. Since oxcarbazepine functions as a prodrug, monitoring focuses on the 10-hydroxycarbazepine metabolite rather than the parent compound [10].
Broad Analytical Ranges for High-Dose Protocols
Mass spectrometry methods accommodate calibration concentrations from 12 ng/mL to 200,000 ng/mL, with top concentrations ranging from 1.5 to 200 μg/mL depending on the analyte [29]. This broad dynamic range supports high-dose treatment protocols while maintaining accuracy within 20% at lowest calibration levels and 15% at higher levels [14]. Correlation factors exceed 0.99 across four orders of magnitude [14].
Adjusting Treatment Based on Genetic Metabolism Profiles
TDM results prompted medication adjustments in 64.2% of clinical visits, most commonly dosage increases occurring in 30.5% of cases [10]. When results indicated subtherapeutic levels, 37.5% received dose escalations, whereas 53.3% showing supratherapeutic levels had dosages decreased [10]. Polymorphisms in CYP3A4, particularly rs2242480, associate with drug responsiveness in pediatric patients, present in 75% of responders versus 43.5% of non-responders [5]. For focal seizures, CT+TT genotypes appeared in 64% of cannabidiol non-responders compared to 33% of responders [5]. Overall, 55.9% of TDM visits resulted in improved outcomes, including reduced seizures in 47.5% and fewer adverse effects in 8.4% [30].
Precision Therapy in Epilepsy: The 2026 Landscape 
Recent molecular discoveries expand precision epilepsy beyond established monogenic disorders into novel genetic territories. SPOUT1 compound heterozygous variants cause autosomal-recessive developmental and epileptic encephalopathy in four unrelated patients, all presenting with infantile epileptic spasms syndrome between three and six months of age [31]. MDN1 variants emerged as epilepsy susceptibility factors in five patients with febrile seizures or secondary epilepsy from the China Epilepsy Gene 1.0 Project [32]. ADGRV1 variants associate with familial febrile seizures and increased sudden unexpected death in epilepsy risk through cardiac and respiratory dysfunction [33] [8].
New Gene Targets: SPOUT1, MDN1, and ADGRV1
Functional analysis using CRISPR-mediated spout1 knockout zebrafish models revealed epileptiform discharges in 14% of knockout fish versus 2% of controls [31]. Transcriptome sequencing demonstrated downregulation of axonal transport genes KIF3A and AP3D1 [31]. ADGRV1 variants were identified in patients experiencing ictal asystole, with in silico analysis predicting protein structure alterations [33].
Antisense Oligonucleotides and Gene Therapy Developments
Antisense oligonucleotides represent the most advanced RNA-based therapeutic modality for precision medicine in epilepsy [13]. Zorevunersen achieved 85% seizure reduction at three months and 73% at six months in Dravet syndrome patients receiving 70 mg doses [34]. Extension studies showed sustained reductions between 58% and 90% over 20 months with 45 mg dosing [34]. ASO treatment for SCN1A increased survival in Dravet syndrome mice by more than fourfold [13]. Single intracerebroventricular injection at postnatal day 2 resulted in 97% survival at 90 days [13].
ASO therapies targeting SCN2A and SCN8A demonstrated similar efficacy in preclinical models. Scn2a ASO administration at postnatal day 1 prolonged median survival from 18 days to 47 days in mouse models [35]. Treatment initiated after symptom onset at day 14-16 also extended lifespan [35]. For KCNA2 mutations, Gapmer ASOs specifically silenced mutated mRNA while preserving normal protein function, restoring potassium channel activity [36].
Machine Learning in Variant Interpretation
Machine learning algorithms reduce variants of uncertain significance across underrepresented populations. Gene-specific algorithms incorporating SpliceAI, AlphaFold protein structures, and gnomAD data achieved positive or negative predictive values exceeding 80% [37]. Among 57,668 epilepsy patients tested, 96% had machine learning evidence applied, with models contributing to benign or likely benign classification in 63% [37]. Asian patients showed 78% definitive classification rates compared to 59% for White individuals [37].
Large language model-assisted frameworks reclassified 60% of variants of uncertain significance, with 24% upgraded to pathogenic and 36% downgraded to benign [38]. These frameworks identified monogenic epilepsy in 36% of reanalyzed cases, oligogenic models in 13%, and modifier variants in 21% [38]. Tools like AlphaMissense, PrimateAI 2.0, and CADD demonstrate accuracy exceeding 0.8 for precision, recall, and F1 scores [39] [40].
Expansion from Rare to Common Epilepsy Syndromes
Genome-wide association studies identify common variants with small individual effects that combine into polygenic risk scores predicting generalized and focal epilepsy risk [4]. Patients with familial epilepsies exhibit higher polygenic risk scores compared to unaffected family members and sporadic cases, regardless of rare variant presence [4]. Despite optimal testing, diagnostic yields reach only 50%, indicating either non-genetic etiologies or incomplete understanding of genetic contributions [4]. Gene therapy approaches divide into gene-specific targeted modulation requiring single gene diagnosis, and broad hyperexcitability modulation independent of specific variants [4]. The latter offers advantages for patients without identified genetic causes, potentially serving broader populations [4].
Conclusion 

Precision epilepsy has progressed from a largely theoretical concept into an increasingly practical component of contemporary neurological care. Advances in molecular genetics, pharmacogenomics, biomarker driven monitoring, and targeted therapeutics have transformed how clinicians understand and manage seizure disorders, particularly in patients whose epilepsy cannot be adequately controlled with conventional antiseizure medications. This shift reflects a broader movement in medicine toward individualized treatment strategies in which therapy is guided by the underlying biological mechanisms of disease rather than by symptom classification alone.
The clinical importance of this transition is substantial. Approximately 30 to 40 percent of individuals with epilepsy continue to experience uncontrolled seizures despite treatment with standard antiseizure drugs. For many of these patients, traditional approaches based on empirical medication selection, seizure type classification, and sequential drug substitution provide limited benefit. Precision epilepsy seeks to address this gap by identifying genetic, metabolic, and pharmacological determinants of treatment response and disease progression, thereby enabling mechanism directed interventions that improve efficacy and reduce avoidable adverse effects.
One of the most established areas of precision epilepsy is pharmacogenomic screening. Genetic testing now plays a critical role in preventing severe drug related hypersensitivity reactions, particularly in populations at increased risk for specific immune mediated adverse events. Screening for susceptibility alleles before initiating certain antiseizure medications has significantly reduced the occurrence of life threatening reactions such as Stevens Johnson syndrome and toxic epidermal necrolysis. This represents a clear example of how genomic information can directly improve treatment safety and influence first line medication selection in routine practice.
Beyond drug safety, precision epilepsy increasingly targets the molecular causes of disease. Gene specific interventions have become especially important in disorders where the pathogenic mechanism is well characterized. In tuberous sclerosis complex, for example, therapies directed at dysregulated mammalian target of rapamycin signaling have provided clinically meaningful seizure reduction while also addressing broader disease manifestations. In glucose transporter type 1 deficiency syndrome, early recognition of impaired cerebral glucose transport allows implementation of ketogenic dietary therapy, which directly compensates for the metabolic defect and often leads to substantial seizure control. These examples illustrate how understanding disease biology can guide highly effective interventions that extend beyond conventional antiseizure pharmacotherapy.
Therapeutic drug monitoring has also evolved within the precision framework. High sensitivity analytical techniques, particularly mass spectrometry based assays, now allow more accurate measurement of antiseizure medication concentrations across diverse clinical contexts. This has improved dosing precision in pediatric patients, pregnant individuals, critically ill patients, and those receiving multiple interacting therapies. By integrating pharmacokinetic data with clinical response and patient specific physiological variables, clinicians can better optimize dosing strategies, minimize toxicity, and maintain therapeutic effectiveness over time.
A particularly important recent development is the emergence of antisense oligonucleotide therapy in genetically defined epilepsies. In disorders such as Dravet syndrome, these therapies are designed to modify gene expression at the RNA level, directly addressing the molecular consequences of pathogenic variants. Early clinical results have shown remarkable therapeutic potential, with sustained seizure reductions exceeding 70 percent in some treated patients. These findings suggest that nucleic acid based therapies may represent a new therapeutic class capable of altering disease trajectory rather than merely suppressing symptoms.
The interpretation of genetic variation has also improved substantially through the integration of machine learning technologies. Advanced computational models now assist in distinguishing pathogenic variants from benign genomic variation, accelerating diagnosis and improving the clinical utility of sequencing data. This is particularly important for underrepresented populations, where historically limited genomic reference datasets have complicated variant interpretation and contributed to diagnostic inequities. Machine learning approaches help refine predictive accuracy by incorporating broader population data, functional evidence, and phenotypic correlations.
Although early precision epilepsy efforts focused primarily on rare monogenic epilepsies, the field is now expanding toward more complex forms of epilepsy with polygenic and multifactorial contributions. Researchers are increasingly investigating how combinations of genetic susceptibility factors, environmental influences, and network level neurobiology interact to shape seizure risk and treatment response in broader patient populations. This expansion is critical because the majority of epilepsy cases do not arise from single gene defects yet may still benefit from biologically informed stratification.
These developments collectively represent a fundamental shift in epilepsy care. Precision epilepsy is redefining seizure management by replacing generalized treatment pathways with interventions tailored to molecular mechanism, metabolic context, and individual pharmacological response. As genomic accessibility improves and targeted therapies continue to emerge, the field is expected to extend its impact beyond specialized centers into broader clinical practice. For neurologists and epilepsy specialists, integrating these advances into routine care offers the opportunity to improve outcomes in patient groups that have historically remained therapeutically underserved.
Frequently Asked Questions: 
FAQs
Q1. What is precision medicine in epilepsy and how does it differ from traditional treatment? Precision medicine in epilepsy uses genetic testing to identify specific molecular causes of seizures and targets them with tailored therapies, rather than using broad-spectrum antiseizure medications that empirically suppress seizure activity. This approach addresses the root molecular defects causing epilepsy, such as metabolic abnormalities requiring specific supplements or dietary interventions, instead of just treating the downstream electrical hyperexcitability.
Q2. How effective is genetic testing in changing epilepsy treatment outcomes? Genetic testing leads to clinical management changes in approximately 50% of tested patients. Among those with follow-up data, about 75% report positive outcomes, with nearly 65% experiencing reduction or elimination of seizures. The diagnostic yield varies by clinical presentation, reaching up to 54% in epilepsy with structural brain malformations and 38% for seizure onset before three years of age.
Q3. Why is HLA-B*15:02 testing important before starting carbamazepine? HLA-B15:02 testing predicts severe cutaneous reactions like Stevens-Johnson syndrome and toxic epidermal necrolysis, which have mortality rates reaching 10-50%. In Han Chinese populations, all patients who developed these severe reactions carried the HLA-B15:02 allele, compared to only 3% of those who tolerated the medication. The FDA mandates screening in genetically at-risk populations before carbamazepine initiation.
Q4. What gene-specific treatments are currently available for epilepsy patients? Several targeted treatments are now in clinical use, including mTOR inhibitors (everolimus) for tuberous sclerosis complex mutations, ketogenic diet for GLUT1 deficiency, pyridoxine for ALDH7A1 variants, perampanel for NR2F1-related epilepsy, and quinidine for KCNT1 mutations. These therapies address specific molecular pathways disrupted by genetic variants rather than broadly suppressing seizure activity.
Q5. How are antisense oligonucleotides changing treatment for Dravet syndrome? Antisense oligonucleotides represent an advanced RNA-based therapy showing remarkable results in Dravet syndrome. Zorevunersen achieved 85% seizure reduction at three months and 73% at six months in treated patients, with sustained reductions between 58-90% over 20 months. This approach specifically targets the genetic cause of Dravet syndrome, offering a mechanism-driven solution for this previously difficult-to-treat condition.
References: 
[2] – https://pubmed.ncbi.nlm.nih.gov/35716052/
[3] – https://practicalneurology.com/diseases-diagnoses/epilepsy-seizures/genetic-mechanisms-of-epilepsy/31588/
[4] – https://pmc.ncbi.nlm.nih.gov/articles/PMC11562134/
[5] – https://www.frontiersin.org/journals/pharmacology/articles/10.3389/
fphar.2025.1626054/full
[6] – https://www.epilepsy.com/causes/genetic
[7] – https://rarediseases.org/rare-diseases/pyridoxine-dependent-epilepsy/
[8] – https://aesnet.org/abstractslisting/identification-of-missense-adgrv1-mutation-as-a-candidate-genetic-cause-of-familial-febrile-seizure-4
[9] – https://pmc.ncbi.nlm.nih.gov/articles/PMC6009246/
[10] – https://pmc.ncbi.nlm.nih.gov/articles/PMC11402380/
[11] – https://www.sciencedirect.com/science/article/pii/S2475299125029658
[12] – https://www.explorationpub.com/Journals/ent/Article/100480
[13] – https://pmc.ncbi.nlm.nih.gov/articles/PMC11551783/
[14] – https://documents.thermofisher.com/TFS-Assets/CMD/Technical-Notes/tn-65139-lc-ms-antiepileptic-drugs-tn65139-en.pdf
[15] – https://pmc.ncbi.nlm.nih.gov/articles/PMC8290505/
[16] – https://www.texaschildrens.org/content/research/fda-approves-everolimus-for-tuberous-sclerosis-associated-seizures-treatment-first
[17] – https://www.thelancet.com/journals/ebiom/article/PIIS2352-3964(25)00184-7/fulltext
[18] – https://pmc.ncbi.nlm.nih.gov/articles/PMC8000344/
[19] – https://www.epilepsy.com/stories/understanding-glucose-transporter-type-1-deficiency-syndrome-glut1-ds-current-management
[20] – https://practicalneurology.com/diseases-diagnoses/epilepsy-seizures/toward-precision-medicine-for-genetic-epilepsies-glucose-transporter-type-1-deficiency-syndrome/32055/
[21] – https://www.seizure-journal.com/article/S1059-1311(16)00023-6/fulltext
[22] – https://www.frontiersin.org/journals/psychiatry/articles/10.3389/
fpsyt.2024.1501238/full
[23] – https://pmc.ncbi.nlm.nih.gov/articles/PMC11960385/
[24] – https://pmc.ncbi.nlm.nih.gov/articles/PMC4158617/
[25] – https://www.frontiersin.org/journals/neurology/articles/10.3389/
fneur.2019.00064/full
[26] – https://www.neurotherapeuticsjournal.org/article/S1878-7479(23)01036-X/fulltext
[27] – https://www.seizure-journal.com/article/S1059-1311(22)00242-4/fulltext
[28] – https://pubmed.ncbi.nlm.nih.gov/38576184/
[29] – https://www.agilent.com/cs/library/applications/5991-8214EN.pdf?srsltid=AfmBOoqXyzMudSnPryV1FHVsIFmYlSQVfyrwEDd0q3ueoJxbvKkTvtbR
[30] – https://www.sciencedirect.com/science/article/pii/S2319417023001178
[31] – https://pmc.ncbi.nlm.nih.gov/articles/PMC11960386/
[32] – https://pmc.ncbi.nlm.nih.gov/articles/PMC12487144/
[33] – https://link.springer.com/article/10.1186/s42494-023-00124-5
[34] – https://www.insideprecisionmedicine.com/topics/precision-medicine/trial-reveals-promising-first-ever-disease-modifying-epilepsy-treatment/
[35] – https://www.jci.org/articles/view/155233
[36] – https://www.regmednet.com/bridging-the-gapmer-promising-gene-therapy-for-rare-epilepsy/
[37] – https://aesnet.org/abstractslisting/the-impact-of-machine-learning-models-in-reducing-variants-of-uncertain-significance-in-individuals-from-underrepresented-populations-who-are-undergoing-genetic-testing-for-epilepsy
[38] – https://aesnet.org/abstractslisting/decoding-the-genetic-complexity-of-epilepsy-an-open-access-ai-llm-powered-variant-reclassification-framework-unraveling-epilepsy-pathogenesis
[39] – https://pmc.ncbi.nlm.nih.gov/articles/PMC11647429/
[40] – https://pmc.ncbi.nlm.nih.gov/articles/PMC12099401/
Video Section
Check out our extensive video library (see channel for our latest videos)
Recent Articles


